Lepbase release 2 went live on 13th February 2016 with 21 annotated assemblies across 17 species.
This release adds six additional species:
- Amyelois transitella v1
- Operophtera brumata v1
- Papilio machaon Pap_ma_1.0
- Papilio polytes Ppol1
- Papilio xuthus Pap_xu_1.0 and Pxut_1.0
- Spodoptera frugiperda v2
Plus updated assemblies/annotations for three of the species in release 1:
- Bicyclus anynana v1.2
- Bombyx mori ASM15162v1 (RefSeq annotations)
- Danaus plexippus v3
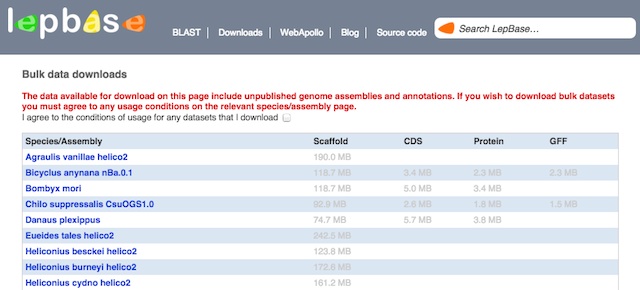
All of these species (together with unannotated Heliconiine DISCOVAR assemblies) are available on our ensembl and blast servers. The files used (assemblies, annotations, protein/cds sequences are available for download, however this includes some unpublished data so please respect the owners of each dataset and contact them to discuss any plans before publishing results based on these unpublished data.
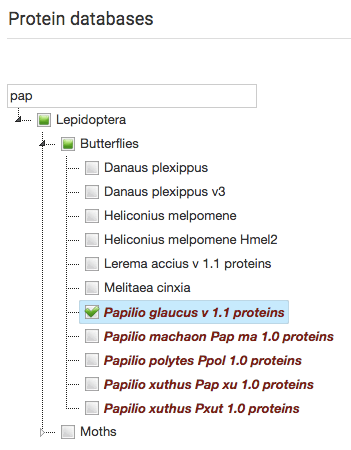
Since release 1 we have also made a few changes to our blast server with a searchable hierarchical listing of blast databases to make it easier to use as the number of available assemblies keeps increasing.
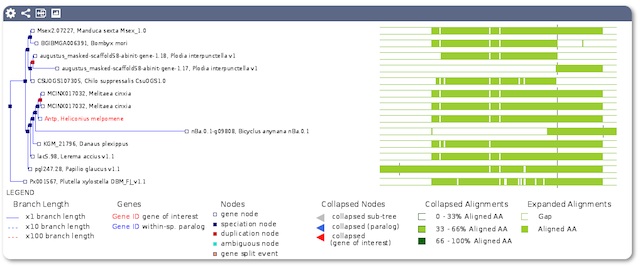
We’ll be focussing on adding more tools and comparative analyses over the next few weeks and months so we’ve also redesigned this website to make it easier to see which tools, analyses and downloads are available.